Tags
anatomically modern humans, David Reich, Denisovans, genetic distance, genus, Neanderthals, race, species, taxonomy
About a year ago I wrote:
The following chart (created by some scientist(s) led by David Reich) shows the genetic divergence between hominin samples as a fraction of the human-chimp difference. So for example, all the human groups have just over a 0.12 genetic divergence with Neanderthals, meaning that the genetic difference between humans and Neanderthals is only 12% as great as the genetic difference between humans and Chimps (source: supplement of Genetic history of an archaic hominin group from Denisova Cave in Siberia.)

The purpose of the chart is to estimate how long ago the different populations diverged from a common ancestor. So since the fossil record tells us that Neanderthals and chimps diverged about 6.5 million years, then humans and Neanderthals should have diverged roughly 0.8 million years ago (12% of 6.5 million) assuming genetic divergence maps to chronological divergence in a linear way.
I transformed the genetic distance matrix into a dendrogram, which looks at all the distances and creates the most parsimonious family tree:
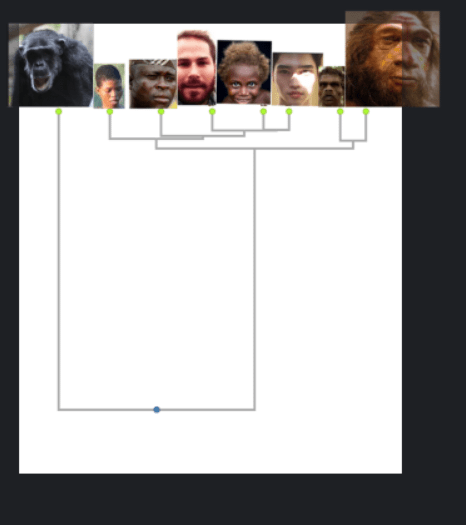
What’s cool about dendrograms is they let you determine the number of categories and subcategories in a very objective way.
Of course dendrograms are only as good as the data you put into them, and I don’t endorse basing taxonomy simply on genetic relatedness, but if I did, here’s how I’d interpret the above tree:
The first major split is between chimps & everyone else. This is consistent with two well recognized genera of hominins : Pan (i.e. chimps) and Homo (humans and near-humans).
Now within the Homo genus, we see another major split in the tree. Anatomically Modern Humans (AMH) vs Archaic Humans. Thus we can divide the homo genus into at least two major species.
Within the Archaic Humans we can further subdivide into major races: Denisovans and Neanderthals.
Now within our own species, AMH, the dendrogram shows three major races: Capoids, Congoids and Non-Africans.
I’m not saying I agree with this taxonomy since it was only based on genetic distance (much of which is junk DNA) but what’s great about using dendrograms is almost everyone looking at them will assign groups to the same categories and subcategories, even if they don’t use the same words (race, species, genus) to describe them. It’s wholly objective.
But what is needed is a dendrogram based on polygenic scores of actual phenotypes. That way people who have the same phenotypes caused by the same genomic architecture could be grouped together.
Unlike the above dendrogram, which groups based on how recently we share a common ancestor, we need to group based on how much of the common ancestor we share.
[update may 26, 2019: an earlier version of this article misspelled dendrogram]
[2nd update may 26, 2019: an earlier version of this article contained bragging that has since been removed]
