Way back in April 2016, I wrote:
Many non-scientists have a great interest in heritability, but lack the science education and/or cognitive ability to understand modern techniques like Genome-wide Complex Trait Analysis (GCTA), so this post is a quick attempt to explain it. Full Disclosure: I have virtually no formal science training beyond high school but this is just an oversimplified explanation.
GCTA gives a measure of the squared correlation between additive genotype and phenotype. The reason it’s so confusing is that you can’t directly correlate a phenotype with a genotype if you haven’t found the genes that code for that phenotype, and thus you can’t determine if someone is genetically high on a given trait.
So for example, you can’t determine if someone’s genetic IQ matches their actual IQ, if you don’t know if they have the genes for IQ. Since a correlation, by definition, is how close the rank order of two variables (i.e. genetic IQ and actual IQ) agree, it can’t be directly calculated if one of said variables (i.e. genetic IQ) can’t be ranked. It would be like trying to calculate the correlation between height and weight, but all the weights were reported in a language you didn’t speak.
To sidestep this problem, GCTA was invented by a scientist of East Asian heritage. In GCTA, instead of ranking everyone in your sample from highest to lowest on each trait, you simply randomly assign people to pairs, and for each pair, calculate the genetic distance and the phenotype distance. So for example, if the people who differ by 100 single nucleotide polymorphisms (SNPs), on average, differ by one standard deviation in IQ, and if people who differ by one standard deviation in IQ differ, on average, by 39 SNPs, then perhaps it can be inferred that (in this sample) the correlation between genetic IQ and actual IQ is whatever number when squared and multiplied by 100, equals 39.
That number is 0.62
This is because in a bivariate normal distribution, the slope of the standardized regression line equals the correlation between two variables, so if a genetic difference of 100 SNPs regresses to a one standard deviation difference in IQ, then one standard deviation must be only 62% as extreme as 100 SNPs and if a one standard deviation difference in IQ regresses to a 39 SNP difference, then 39 must be only 62% as extreme as one standard deviation.
Once we have the correlation of say 0.62 between additive genotype and phenotype , we square it to get the amount of variation explained which in this example would be 0.38 (the real number is probably much higher, and even higher still for broad-sense heritability).
Of course what very few people realize is that heritability is technically NOT the percentage of the phenotypic variation explained by genes, it’s the percentage explained by genes when environment is held constant or allowed to vary randomly.
Recently commenter Trumpocalypse (aka Mugabe), also commented on GCTA:
if it could be shown that the joint distribution, f, of some measure of genetic distance between individuals, d, and phenotypic difference, p, has a linear conditional mean (that is, f(d|p) is a straight line), then it would be possible to avoid all the problems of twin studies and the shared womb of twins…
the conditional mean could simply be extrapolated to d = 0. and from the mean difference, p, at d = 0, the heritability could be calculated easily.
furthermore the perfect-ness of the measure of genetic distance would be irrelevant. the joint probability density function would “include” this error. so with a large enough sample the extrapolation would have very little error.
this is essentially what GCTA is.
but whether the joint distribution is bivariate normal or some other distribution with linear conditional mean…i don’t know.
Here is some genetic distance data on nine major human populations:
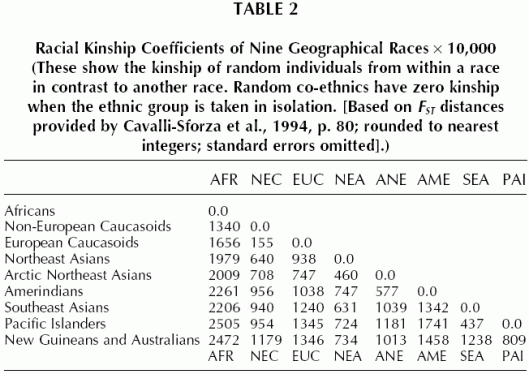
According to Richard Lynn’s controversial meta-analysis of these nine genetic clusters, Africans average IQ 67, Non-European Caucasoids average 84, European Caucasoids average 99, Northeast Asians average 105, Arctic Northeast Asians average 91, Amerindians average 86, Southeast Asians average 87, Pacific Islanders average 85, and New Guineans and Australians average 62.
Although this is group data, not individual data, it would be interesting to compare group genetic distance to group IQ difference. If for example, I knew both the average IQ difference between two random humans from anywhere in the World, and if I also knew the Fst distance between two random individuals from anywhere in the World, I think I could probably use my crude understanding of GCTA, to estimate the IQ phenotype-genotype individual level correlation of the entire human species from Sforza’s and Lynn’s group level data. Squaring this correlation might be a good proxy for IQ’s Worldwide heritability.
On the other hand, the fact that Sforza intentionally tried to use non-selected genes to calculate genetic distance (since these mutate at a regular rate creating a reliable molecular clock for splitting off dates) might make the exercise pointless.

1. Can gcta be used on relatives of the same race to determine iq reasonably? How about mixed race individuals?
2. Can gcta be used with different generations to find iq?
3. Does convergent evolution for intelligence lead to similar nucleotides?
4. For gcta, do you think BRCA gene sequence be used? On a similar note, do the studied DNA sequences have anything to do with iq?
Not sure i understand your questions. GCTA can’t determine anyone’s IQ, it can only determine how much of the variance in IQ is narrowly genetic
broadly genetic if it satisfies the criteria i gave.
all that’s required is a robust statistical relationship be demonstrated.
One could certainly establish a statistical relationship between genetic distance and IQ difference using Cavalli-Sforza & Lynn’s data, but it would be a group level relationship, not an individual relationship.
yes. surprising for HBDers is that two individuals from different races can be closer genetically than two individuals from the same race.
naively HBDers think that the genetic distance between individuals within any racial group is almost always much smaller than that between individuals from two different races. that is, first they categorize people by race, then by other characteristics. race is first for them formally, in their categorizing schema, so they naively assume it is also first genetically.
but this is not the case.
race is defined by a small subset of human genes. maybe < 5%. the rest of the genes vary willy-nilly.
a japanese and an ituri forest pygmy may be more alike genetically than either is to a random member of their own group.
members of the same race are obviously much closer in terms of their most recent common ancestor, but humans are such a new species that this identity by descent carries little information about an individual's genome.
humans are so homogeneous in genetic terms, despite their sometimes striking difference in appearance, that…
the genetic variation within a troop of chimps, man's closest ancestor genetically, is greater than that of a swede and an abo.
humans look very different superficially, but all humans speak. all humans can learn to speak the king's english, including abos.
from the pov of an alien, humans are like thoroughbred race horses…
very little genetic variation.
so from the pov of an alien, the likelihood that selective breeding (or CRISPR gene-editing) of humans for IQ will produce people smarter than any human has ever been is the same as the likelihood that there will ever be a horse faster than secretariat…
NIL!
secretariat's 1973 belmont is another finger in the eye of HBD.
Swank argued that the human-chimp IQ gap was only 60 points and took many millions of years to evolve, so how could even a 15 point IQ gap evolve in the last 50,000 years, let alone 800 (Cochran’s Ashkenazi theory), unless the high IQ group had extemely small variance as a result (compared to the low IQ group)since high IQ mutations must be incredibly rare
I responded by arguing the human-chimp gap was actually hundreds of points, but inter-species IQ comparisons are admittedly pseudoscience at this point
Cochran’s Ashkenazi theory is the definition of a just-so story.
btw, given the fall in the loonie i bought shares in a canadian bank today.
Canadian Western Bank.
presumably, you can do the same with the European data of genetic distance..
off topic, I was thinking about that genetic distance data the other day. It occurred to me that there must be a lot of genetic distance across time. That is, modern Italians compared to italians 2000 years ago. How large would that genetic distance be? Would it be comparable to the distance between the english and the russian today? or bigger?
much much smaller.
italian is much more like latin than latin was like old slavic or old german.
the slavic languages differentiated later than the germanic languages.
polish, czech, serbo-croatian, etc. were all the same language 1000 years ago.
whereas old english, old norse, and old german were already separate languages 1000 years ago, as were french, spanish, and italian.
and that’s a huge problem for HBDers.
contemporary…
iranians = ancient persians.
tunisians = carthaginians.
lebanese = phoenicians.
italians = romans.
greeks = greeks.
iraqis = babylonians.
syrians = syrians.
germans = human sacrifice practicing filthy illiterate savages.
BUT
ashkenazi jews != ancient jews.
such jews have a lot of italian and other european admixture.
i guarantee no ancient jew looked like yitzhak rabin.
That’s why I think genetic distance using neutral genes is a silly way to define race, beause those genes are used precisely because they don’t tell us how similar people are, only how long ago they separated
Sub-Saharans & Papuan New Guineans are virtually indistinguishable, yet in terms of the genes Cavalli-Sforza measured, both groups are closer to the British than they are to each other
Genetic distance is a misnomer. It’s actually chronological distance as measured by genes.
What people don’t get is any group of people that diverged from Italians less than 2000 years ago would be considered more similar to modern Italians than the ancient Italians were, even if they evolved into worms in that time
Because all it’s measuring is time.
the comment lion won’t post on Game of Thrones:
is this a joke?
it’s so horrible i couldn’t take more than 10 minutes of it.
puh
puh puh
prole.
I’m stunned that he thinks it’s the best series ever. It’s not even the best HBO series ever. Has he not seen Six Feet Under or The Sopranos?
the only shows i watch, but only via internet piracy are:
1. South Park.
2. Mike Tyson Mysteries.
3. Anthony Bourdain’s No Reservations.
i find all drama and fiction to be tedious and tiresome and stupid. and the tv or movie versions are even worse. the bbc hasn’t had a good series since Brideshead. and that was 1981 iirc.
i don’t own a tv.
but there are a few good movies and novels…
a very very few.
Movies used to be considered artistically superior to TV, but that perception has changed because of cable channels like HBO
oh wait bourdain’s new show is called Parts Unknown.
whatever.
a little appreciated effect of mass media is…
mass media is not only propaganda. it is also a distraction. the masses can be controlled in two ways.
1. they can be convinced of the un-truth which serves their masters.
2. they can be distracted perpetually. their heads can be stuffed with trivia such that they lose whatever bearings the may have had. they are permanently infantilized.
and now movie stars are doing tv.
media is a very bad investment, and will be thenceforth and forever.
the real issue is internet piracy.
gentiles ripping off jews.
praise be to be allah.
what?
i don’t know man?
despite the brain damage, mike tyson is far more impressive intellectually than ali or the klitschkos.
and no fighter has ever been as dominant in his prime.
tyson is the capablanca of boxers.
My favourite TV series was called Cracker and was about a psychologist (Robbie Coltraine) working with the Manchester police – it is quite simply brilliant!
an IQ test for humans.
an IQ test for neanderthals.
an IQ test for rats.
my own subjective experience is that what “the laity” mean when they say that so-and-so is smart or that so-and-so is smarter than so-and-so is…
much more strongly correlated with the VIQ…but not perfectly.
this is even true of dogs, though again not perfectly.
that is, a dog’s intelligence is largely appraised by his ability to respond to verbal cues.
my own, perhaps biased thinking is…
the wechslers may weight the verbal too little.
the SB and SAT, etc. may weight the verbal too much.
and the reason is very simple:
the verbal tests are power tests. you get it or you don’t.
the performance tests are largely speed tests. everyone gets it, just some faster than others.
in dog terms…
the airdale terrier is not an especially fast learner.
but once he learns something, he never lets it go.
the airdale is what i mean by smart.
if IQ were really “like height”, then those who continue to develop (intellectually) throughout their lives…however slowly…
these are the smart people…
eventually.
i recall a guy i went to school with who went on to earn a masters in mechanical engineering from berkeley.
he made higher SATs than i did, but one day…
i was in the 7th grade, he in the 8th…
he had enormous trouble doing mazes and he couldn’t play chess worth a tinker’s damn.
i mean it wasn’t just that he hadn’t ever played chess or done a maze.
my impression was…
wtf? it’s not that hard. what’s wrong with you?
at the same time, peepee will be happy (or unhappy) to know that…
the one guy who all the smart guys in my hs thought was even smarter than them…
he was the one guy who went to harvard.
last i could tell he was working in london in finance.
and that guy didn’t only have a higher VIQ than me.
he was a better chess player.
but his brother was a “bone head”. he said so himself.
so much for h^2.
and he was a red haired freckled goy.
and he was a red haired freckled goy.
If East Asian supremacism doesn’t work out, i may forget about race & focus instead on just color
The lighter the hair, eyes & skin, the smarter on average
After all Bill Gates is the World’s richest man & blue eyes & white skin appeared very late in evolution
What it sounds to me like you are describing has been done:
http://www.sciencedirect.com/science/article/pii/S019188691630174X
Also, you don’t square the correlation when you calculate heritability. This is counter-intuitive to people who are used to statistics in general but not especially behavioral genetics. Jensen authored a widely read paper explaining this in the early 1970’s:
http://psycnet.apa.org/index.cfm?fa=buy.optionToBuy&id=1971-09170-001
right.
h^2 is…
the correlation between the scores of MZAs.
BUT…
if one had the score of one MZ twin, the score of the other would have SD =
sqrt(1 – (h^2)^2).
two squares!
the score of the other would, theoretically, have a normal distribution with mean (h^2)*(IQ of twin) and stdev of sqrt(1 – (h^2)^2).
And the stains comin’ from my blood tell me, “Go back home.”
You square the genotype-phenotype correlation (ideally among adopted people) to get heritability. The correlation between identical twins raised apart is not the genotype-phenotype correlation, it’s the phenotype-phenotype correlation of people with identical genotypes, hence it’s the SQUARED genotype-phenotype correlation.
exactly.
Yea are correct. Sorry for the misunderstanding, I only skimmed this post and made an incorrect assumption about what you meant.
Yea are correct. Sorry for the misunderstanding
It’s all good. Thanks for the link you provided.
Wow the world must be ending; Mugabe just agreed with PP.
the world ended a long time ago.
OMG it has been done. And so recently. Weird.
what’s been done?
What’s been done?
See the link Sean Last provided
“Chances and limits of this approach (e.g. no intelligence coding genes detected)…”
Weird…
But they didn’t calculate worldwide heritability. Only the correlation between genetic distance and IQ difference, at the group level, impressively with environment controlled (to the extent it can be)
seriously peepee…
buy CWB on margin.
is that allowed in canada?
2.5x margin is allowed in the US.
do it!
Canadian Western Bank!
the best popular de-construction of peepee-ism.
that i know of…
after the money’s gone…
a less popular repudiation of peepee-ism:
the ne plus ultra…
if you don’t “get it” by watching this…you can’t get it.
monkery is faggotry…
obviously…
with one exception…
the 300 or so carthusian monks (and ??? nuns)…
they ARE…
ARE…
the UPGRAYEDDs of the world.
how many people are selling???
you can be “successful” if only you…
…
to be “successful” in the contemporary world means that one is esteemed by men…not invisible.
…
all carthusians have unmarked graves.
should any publish something in his lifetime, its author is “a carthusian”…not the name of the monk.
but the hoi polloi…
they imagine that…
their fame…
should they ever achieve such…
it will preserve them…
IT WON’T!
SIC TRANSIT GLORIA MUNDI.
…
I met a traveller from an antique land
Who said: “Two vast and trunkless legs of stone
Stand in the desert . . . Near them, on the sand,
Half sunk, a shattered visage lies, whose frown,
And wrinkled lip, and sneer of cold command,
Tell that its sculptor well those passions read
Which yet survive, stamped on these lifeless things,
The hand that mocked them, and the heart that fed:
And on the pedestal these words appear:
‘My name is Ozymandias, king of kings:
Look on my works, ye Mighty, and despair!’
Nothing beside remains. Round the decay
Of that colossal wreck, boundless and bare
The lone and level sands stretch far away.”
THERE IS WATER AT THE BOTTOM OF THE OCEAN!
Btw Mugabe, it seems the Raven might be more heritable than you’ve claimed:
http://genetictwinstudies.weebly.com/uploads/1/5/4/5/15451376/2194164.png?423
http://genetictwinstudies.weebly.com/minnesota-twin-study.html
i never claimed anything regarding the RPM’s h^2.
what i claimed was that the RPM correlates poorly with other self-described IQ tests.
and this is…
TRUE!
here we are now. entertain us.
alekhine beat capablanca…
pronounced…
AL..
YEA…
KEEN…
and what was the most salient difference between the two?
AL-YEA-KEEN was an
AL-
KOH-
HALL-
ICK.
yea!
Hey, yea!
I’m worse at what I do best…
as has been pointed out by marion in an obscure (french) way…
there are many reasons why people may contemn some proposition…
think of them.
all of them.
all of the reasons why people may contemn some proposition.
quodlibets included…
kerouac puts his alcoholic finger over his eye…
he died from cirrhosis age 47…
marion said…
sometimes…
the opposition to whatever….
is motivated by…
UN-Cynicism.
that is…
sometimes…
sometimes…
the opposition to a given proposition is…
that…
IT IS THE TRUTH!
kerouac is an interesting character….
why?
it’s gay to say…
but it’s true…
he was very good looking.
his interviewer, steve allen, said the same…
a guy who looked like a movie star but was “shy”.
kerouac is also interesting as…
either he had very narrow shoulders or he had a ginormous head.
and…
he might be mistaken for an idiot…
but he wasn’t…